炎症性遺伝子シグネチャー
炎症性遺伝子シグネチャーとは、特定のタイプの遺伝子発現プロファイル(GEP)であり、腫瘍微小環境の炎症シグネチャーを評価するために使用されます。これらはがんの種類によって異なり、強力な診断ツールとなる可能性があります。
がんバイオマーカーとしての妥当性
- 特定のタイプのGEPである炎症性遺伝子シグネチャーは、腫瘍微小環境の炎症シグネチャーを評価するために使用されます1。
- GEPにより、膨大な遺伝子群のmRNAの発現状況が評価されます2。
- これにより、明確な分子プロファイル(または、遺伝子シグネチャー)が形成され、細胞機能の全体像が得られま
す2,3。
- 炎症過程に関与する、特定遺伝子の発現を組み合わせた炎症性遺伝子シグネチャーにより、腫瘍微小環境内における免疫細胞の存在が示される可能性があります1,3。 - 炎症性遺伝子シグネチャーは、がんの種類によって異なり、強力な診断ツールとなる可能性があります3-6。
- こうした遺伝子シグネチャーは、腫瘍微小環境における免疫活性の代替指標(surrogate marker)として使用できる可能性があります4。 - インターフェロンγ(IFN-γ)遺伝子の発現は、免疫応答の活性化に重要な役割を果たしています7。
- IFN-γ等の炎症性遺伝子シグネチャーを用いて、腫瘍微小環境内の炎症の程度を評価することができます8。
炎症性遺伝子シグネチャーは、潜在的な効果予測I-Oバイオマーカーとして研究されています。
評価法
- 遺伝子シグネチャーを評価する際、複数遺伝子の発現レベルを決定するための媒介指標として、mRNAの発現状況が使用されます2。
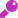
- 次世代シーケンサー(NGS)を用いたRNA seqおよび遺伝子発現パネルにより、GEPの評価が可能となります(遺伝子発現プロファイルは下記のように示されます。ご参照ください)9,10。
Gene Expression Panel
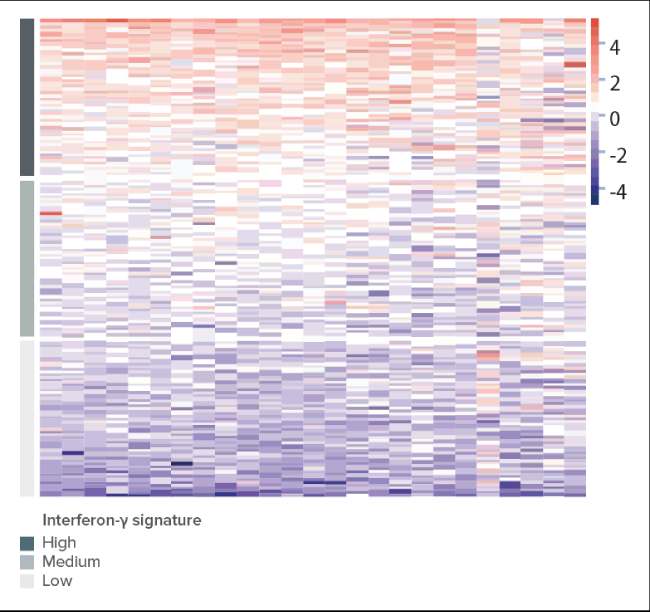
REFERENCES–炎症性遺伝子シグネチャー
- Torri A, Beretta O, Ranghetti A, et al. Gene expression profiles identify inflammatory signatures in dendritic cells. PLoS One. 2010. doi:10.1371/journal.pone.0009404.
- Walker MS, Hughes TA. Messenger RNA expression profiling using DNA microarray technology: diagnostic tool, scientific analysis or un-interpretable data? (review). Int J Mol Med. 2008;21(1):13-17.
- Linsley PS, Chaussabel D, Speake C. The relationship of immune cell signatures to patient survival varies within and between tumor types. PLoS One. 2015. doi:10.1371/journal.pone.0138726.
- Galon J, Angell HK, Bedognetti D, Marincola FM. The continuum of cancer immunosurveillance: prognostic, predictive, and mechanistic signatures. Immunity. 2013;39(1):11-26.
- Gibney GT, Weiner LM, Atkins MB. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 2016;17(12):e542-e551.
- Cesano A, Warren S. Bringing the next generation of immune-oncology biomarkers to the clinic. Biomedicines. 2018. doi:10.3390/biomedicines6010014.
- Ikeda H, Old LJ, Schreiber RD. The roles of IFNγ in protection against tumor development and cancer immunoediting. Cytokine Growth Factor Rev. 2002;13(2):95-109.
- Abiko K, Matsumura N, Hamanishi J, et al. IFN-γ from lymphocytes induces PD-L1 expression and promotes progression of ovarian cancer. Br J Cancer. 2015;112(9):1501-1509.
- Fumagalli D, Blanchet-Cohen A, Brown D, et al. Transfer of clinically relevant gene expression signatures in breast cancer: from Affymetrix microarray to Illumina RNA-Sequencing technology. BMC Genomics. 2014. doi:10.1186/1471-2164-15-1008.
- Cesano A. nCounter® PanCancer Immune Profiling Panel (NanoString Technologies, Inc., Seattle, WA). J Immunother Cancer. 2015. doi:10.1186/s40425-015-0088-7.
- Finotello F, Di Camillo B. Measuring differential gene expression with RNA-seq: challenges and strategies for data analysis. Brief Funct Genomics. 2015;14(2):130-142. 12.Sharma P, Retz M, Siefker-Radtke A, et al. Nivolumab in metastatic urothelial carcinoma after platinum therapy (CheckMate 275): a multicentre, single-arm, phase 2 trial. Lancet Oncology. 2017;18(3):312-322.
















